In 2005, Ray Kurzweil wrote a book about “The Singularity” which is when machine intelligence would surpass human intelligence and potentially merge with it. This prediction was grounded in the exponential growth of computing power that has persisted since the earliest electronic computers of the 1930s.. It is entertaining to read how some readers disagree with this interpretation, but it is also striking that one of the predictions in the book actually happened four years early.
While Kurzweil famously predicted that a machine would pass the Turing Test by 2029, conversational AI systems like ChatGPT arguably reached this milestone in 2025.
Many things would need to happen in order for Kurzweil’s nerd apotheosis to arise in twenty years. While the desirability of a human-machine fusion may be a fascinating topic for a future post, today it is interesting to consider other ‘singularities’ that could develop sooner than 2045. As we wait for the future to decide whether it needs us, let’s focus on a narrower question that nevertheless has significant implications.
Early Attempts at Solving the Protein Folding Problem
The protein folding problem, predicting a protein’s three-dimensional structure from its amino acid sequence, has long been one of the grand challenges of molecular biology. The earliest publication I have found on the application of neural networks to the protein folding problem was published in 1988 by Qian and Sejnowski, in a highly innovative paper. They used a neural network to predict the secondary structure of proteins and achieved a dismal (by today’s standards) accuracy of 64%. This state of affairs continued into the 2000s, and predicting a tertiary structure to any useful accuracy was essentially impossible unless a closely related protein’s structure was known through painstaking structural biology work.
For decades afterward, meaningful tertiary structure prediction remained nearly impossible unless a closely related protein structure had already been experimentally determined through labor-intensive methods like X-ray crystallography or NMR spectroscopy.
AlphaFold and the AI Breakthrough in Protein Structure Prediction
But in 2018 neural networks and large protein structure datasets built over decades were brought together to achieve what amounts to a solution of the protein folding problem. Given arbitrary sequences of amino acids, Google DeepMind’s AlphaFold now reliably predicts protein structures that are nearly as good as atomic structures determined by biophysical methods (table 1). Powerful GPUs combined with large training sets of validated data made this achievement possible. The significance of this breakthrough was underscored when key contributors to this work were awarded the 2024 Nobel Prize in Chemistry.

From Protein Prediction to Protein Design
Closely following protein structure prediction came something even more powerful: computational protein design.. Not only were we now able to predict a structure from its sequence, but the reverse problem of finding a sequence that would fold into a particular structure was beginning to yield to related computational tools. The program RF Diffusion, first described in 2023, uses generative AI methods to design protein backbones from scratch. In combination with ProteinMPNN which very efficiently finds amino acid sequences that fit any protein backbone structure, it is now possible to design proteins more efficiently than ever before. Prof. David Baker shared the Chemistry Nobel Prize in 2024 for his contributions to to computational protein design. Computational biologists are releasing new computational tools for protein design at a frenetic pace and while a recent review gives a nearly comprehensive account of available design software, further developments could already justify updating the review.
AI-Designed Protein Binders: Beyond Traditional Antibodies
Once you have the ability to design proteins, some applications become readily apparent. One of these is the task of generating ‘protein binders’, proteins that bind target proteins of interest. While antibodies have been used for decades as tools to recognize, detect, and purify specific proteins, designed binders are now gaining recognition as useful tools that could replace antibodies produced by immunizing animals.
In 2024, I co-founded with two colleagues a company called Alces Bioworks with the goal of using protein design to generate binders we call Alceins. These designed proteins can be used as affinity reagents for assays such as ELISAs, Western blots, immunohistochemistry, immunofluorescence, protein detection and purification, much like antibodies have been used in the past. Alceins can be designed and tested in a few weeks instead of several months like antibodies and, because they are designed, their structure is known. This offers news possibilities in life sciences R&D and manufacturing while lowering costs and reducing the use of animals.
And proteins binding to other proteins is just one function, albeit a crucial one, that proteins perform. They also bind to a variety of other molecules with great affinity and specificity, such as long polymeric nucleic acids like DNA and RNA, and smaller molecules of all kinds. But perhaps the most remarkable and critical function of proteins is catalysis. The ability of certain proteins called enzymes to accelerate an almost infinite diversity of chemical reactions is one of the most striking capabilities of proteins, and it is fundamental to life. In fact, enzymatic catalysis can be thought of as a special case of protein binding wherein an enzyme doesn’t merely bind the substrate(s) of a reaction, but it also binds and stabilizes the transition state of the reaction to greatly accelerate its progression to completion.
While early efforts at protein design required the synthesis, expression, and screening of tens of thousands of genes (see Table 2), it is now possible to make 10 to 100 designs and have a reasonable expectation of finding a binder with sub-micromolar affinity. This increased efficiency means that it takes much less time and effort to find binders for arbitrary targets of interest. However, the experimental step of identifying a binder, even from a small collection of designs, is not yet trivial, requiring the synthesis of multiple genes, their expression, and the purification of the resulting proteins in order to test their binding affinity to the target of interest. That’s exactly what we do at Alces.

Defining the Protein Design Singularity
Which brings us to the Protein Design Singularity (PDS). We define the Protein Design Singularity as the moment when designing a protein with a specified function becomes 95% efficient.
In practical terms, 19 out of 20 AI-generated protein designs will work exactly as intended. At that point, biology transitions from a trial-and-error science to a programmable engineering discipline.
What Happens After the Protein Design Singularity?
The Protein Design Singularity will be significant because of the efficiency it will bring to the life sciences. Biology would undergo a phase transition, from a trial-and-error discipline to a truly programmable endeavor. No longer will we need to screen dozens of designs to arrive at one functional novel protein. It will be possible to design multi-subunit protein complexes, sophisticated molecular machines that will be used to detect target molecules and process them in new ways. We could design ‘teams’ of proteins that carry out signaling inside our cells to diagnose and treat disease. Logical operations could be carried out in engineered microorganisms to perform complex behaviors such as degrading microplastics in response to predetermined cues.
The Protein Design Singularity would herald an era of unprecedented control over our world of molecules. And when will this Singularity happen?
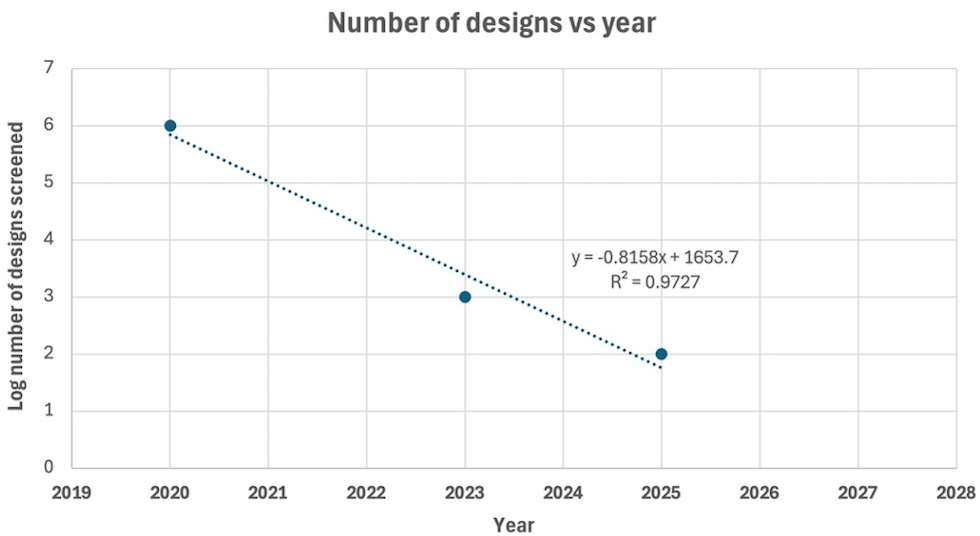
When Will the Protein Design Singularity Arrive?
According to the above graph, it would be February 1st, 2027. About one year from now. Of course, this analysis has only three data points and is therefore lacking in precision, and it could be debated whether this curve is as log-linear as shown here. But the general trend has been quite striking. Fewer and fewer designs need to be screened to arrive at a functional ‘binder’. Exciting new capabilities in biotechnology seem to be quite near.
Conclusion: A New Era for Biotechnology
The convergence of AI, protein engineering, and life sciences is creating capabilities once thought to be science fiction. Just as AlphaFold solved protein folding, protein design is rapidly approaching a singularity of its own.
When that happens, our ability to engineer the molecular foundations of life will redefine the life sciences.
In the future, new biology will not only be discovered, it will be designed.
FAQs
The Protein Design Singularity (PDS) is the proposed point at which proteins can be designed with approximately 95% probability of success. Not merely that they express and fold as intended, but that they also do so exactly as intended. At that point, protein design shifts from trial-and-error screening to a programmable, engineering-driven discipline. This milestone would dramatically accelerate biotechnology research, drug discovery, and synthetic biology applications.
AlphaFold, developed by Google DeepMind, revolutionized biology by accurately predicting a protein’s three-dimensional structure from its amino acid sequence. Introduced in 2018, AlphaFold achieved near-experimental accuracy, effectively solving the decades-old protein folding problem. This breakthrough significantly reduced reliance on time-intensive methods like X-ray crystallography and NMR spectroscopy and laid the foundation for modern computational protein design.
Protein structure prediction determines how a given amino acid sequence folds into a three-dimensional structure. Computational protein design reverses this process by identifying amino acid sequences that will fold into a desired structure or perform a specific function. Tools such as RF Diffusion and ProteinMPNN use generative AI and machine learning to design novel protein backbones and sequences, enabling the rapid creation of protein binders, enzymes, and molecular machines.
AI-designed protein binders are emerging as superior alternatives to traditional antibodies. Companies like Alces Bioworks design custom protein binders (Alceins™) in weeks rather than months, with known molecular structures and predictable binding properties. These engineered binders can be used in ELISA, western blotting, immunofluorescence, and other life sciences assays. As computational protein design efficiency improves, AI-generated binders will increasingly replace animal-derived antibodies in research and biomanufacturing.
